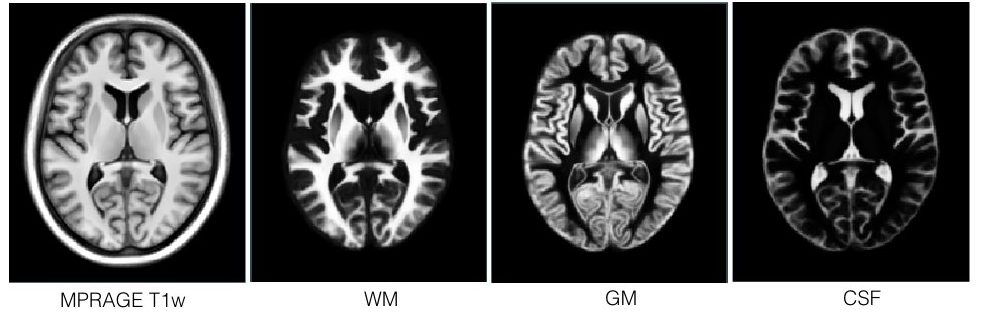
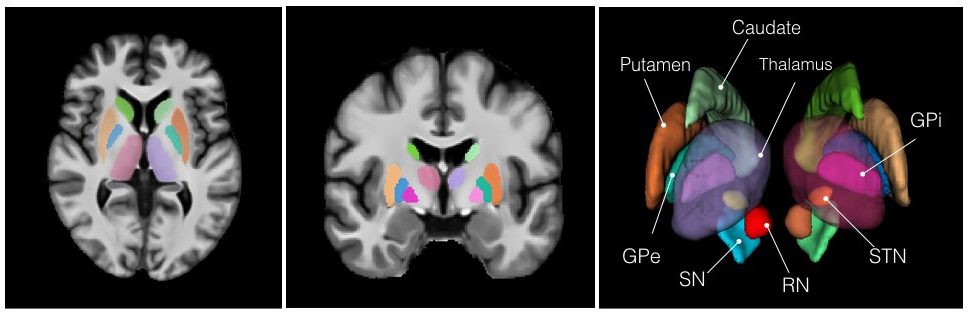
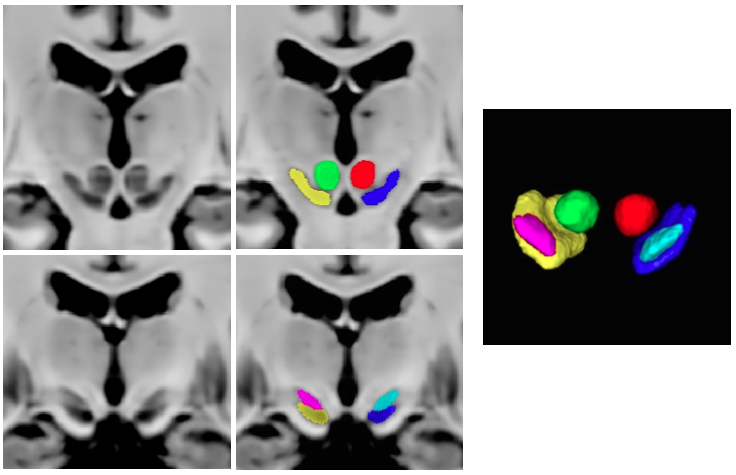
T1-T2* fusion: PD25-fusion-template-{0.3mm, 0.5mm, 1mm}
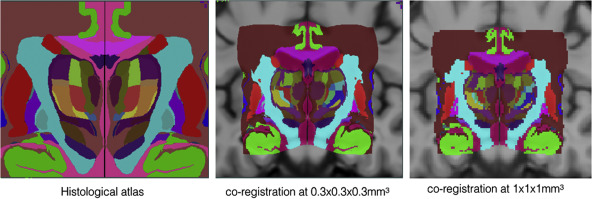
Co-registered histological atlas: PD25-histo-{0.3mm, 1mm}
Histological labels: PD25-histo-labels.csv
BigBrain co-registration
1. Deformed BigBrain atlases:
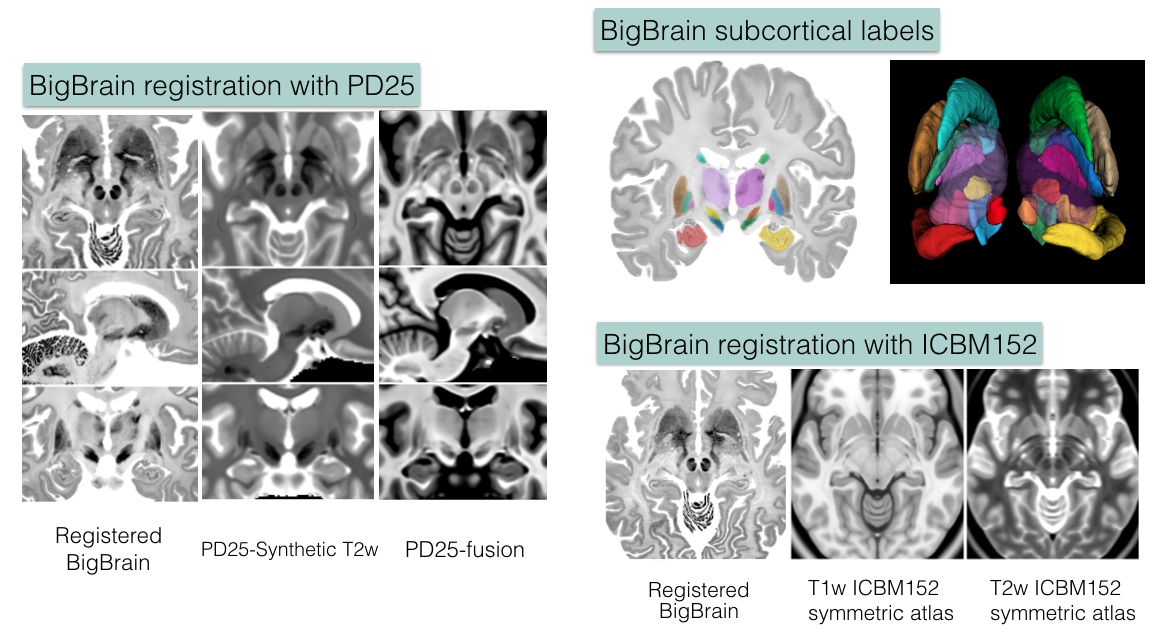
- BigBrain in PD25 space: BigBrain-to-PD25-nonlin-{300um, 0.5mm, 1mm}
- BigBrain in ICBM152 symmetric atlas: BigBrain-to-ICBM2009sym-nonlin-{300um, 0.5mm, 1mm}
- BigBrain in ICBM152 asymmetric atlas: BigBrain-to-ICBM2009asym-nonlin-{300um, 0.5mm, 1mm}
- Synthetic T2w PD25 atlas: PD25-SynT2-template-{300um, 0.5mm, 1mm}
- T1-T2* fusion PD25 atlas: PD25-enhanceFusion-template-{300um, 0.5mm, 1mm}
2. Manual subcortical segmentations:
- BigBrain coregistered to ICBM in the BigBrain2015 release: BigBrain-segmentation-0.3mm
- MNI PD25: PD25-segmentation-0.5mm
- ICBM152 2009b symmetric: ICBM2009b_sym-segmentation-0.5mm
- ICMB152 2009b asymmetric: ICBM2009b_asym-segmentation-0.5mm
3. Related transformations:
- BiBrain-to-PD25: BigBrain-to-PD25-nonlin.xfm
- BigBrain-to-ICBM2009asym: BigBrain-to-ICBM2009asym-nonlin.xfm
- BigBrain-to-ICBM2009sym: BigBrain-to-ICBM2009sym-nonlin.xfm
- PD25-to-ICBM2009asym: PD25-to-ICBM2009asym-nonlin.xfm
- PD25-to-ICBM2009sym: PD25-to-ICBM2009sym-nonlin.xfm
4. List of subcortical labels: subcortical-labels.csv
Publications
- Y. Xiao, V. Fonov, S. Beriault, F.A. Subaie, M.M. Chakravarty, A.F. Sadikot, G. Bruce Pike, and D. Louis Collins, “A dataset of multi-contrast population-averaged brain MRI atlases of a Parkinson’s disease cohort,” accepted in Data in Brief, 2017.
- Y. Xiao, V. Fonov, S. Beriault, F.A. Subaie, M.M. Chakravarty, A.F. Sadikot, G. Bruce Pike, and D. Louis Collins, “Multi-contrast unbiased MRI atlas of a Parkinson’s disease population,” International Journal of Computer-Assisted Radiology and Surgery, vol. 10(3), pp. 329-341, 2015.
- Y. Xiao, S. Beriault, G. Bruce Pike, and D. Louis Collins, “Multicontrast multiecho FLASH MRI for targeting the subthalamic nucleus,” Magnetic Resonance Imaging, vol. 30, pp. 627-640, 2012.
If you are using the BigBrain atlas co-registration dataset, please refer to the following preprint:
- Y. Xiao, J.C. Lau, T. Anderson, J. DeKraker, D. Louis Collins, T. Peters, and A.R. Khan, “Bridging micro and macro: accurate registration of the BigBrain dataset with the MNI PD25 and ICBM152 atlases,”
If you are using the Big Brain data, please cite the following publication:
- Amunts, K. et al.: “BigBrain: An Ultrahigh-Resolution 3D Human Brain Model”, Science (2013) 340 no. 6139 1472-1475, June 2013
Copyright
Copyright (C) 2016,2017,2018 Yiming Xiao, McConnell Brain Imaging Centre,
Montreal Neurological Institute, McGill University.
License
PD25 atlases are distributed under CC BY-NC-SA 3.0 Licence
Dataset for BigBrain co-registration with PD25 and ICBM152 is under CC BY 4.0 Licence. Note that this exception to the existing BigBrain dataset does not alter the general term of the license for the use of BigBrain itself, which is still under CC BY-NC-SA 4.0 License.
Download
Version 20170213: Download archives containing brain atlases, brain masks, midbrain and subcortical segmentation and histological labels: MINC1, MINC2, NIFTI
Version 20160706: Download archives containing brain atlases, brain masks and midbrain segmentation: MINC1, MINC2, NIFTI
Co-registration of BigBrain with PD25 and ICBM152 atlases: Download archives containing registered Big Brain atlas, manual segmentations, and registration transformation (only available in MINC2 package): MINC2, NIFTI